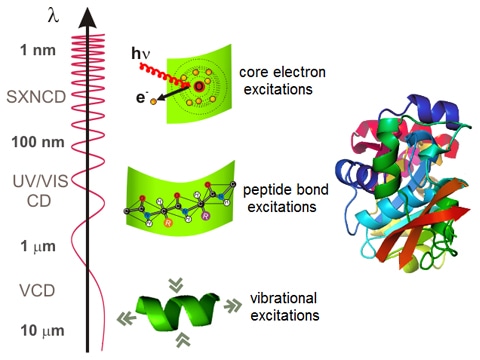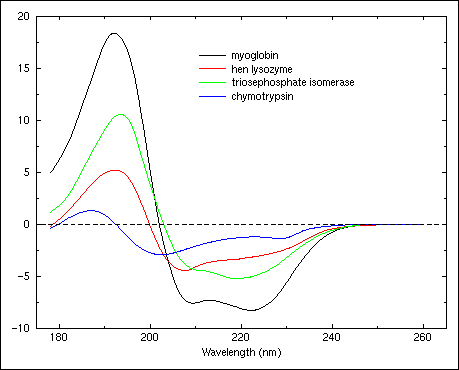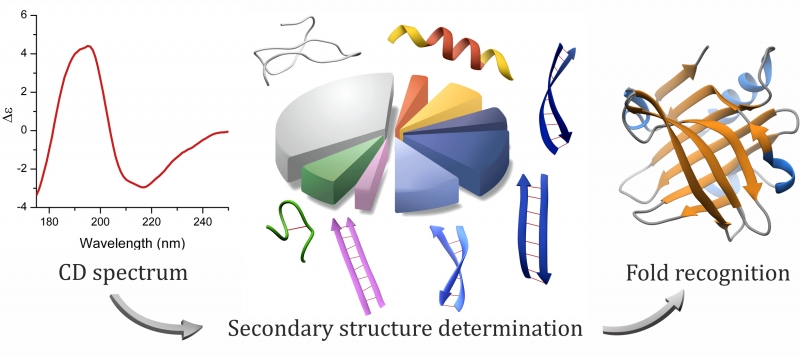
A new method to identify protein secondary structure and fold | French national synchrotron facility

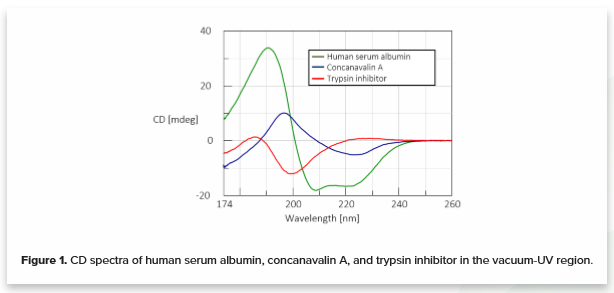
SESCA: Predicting Circular Dichroism Spectra from Protein Molecular Structures | Journal of Chemical Theory and Computation
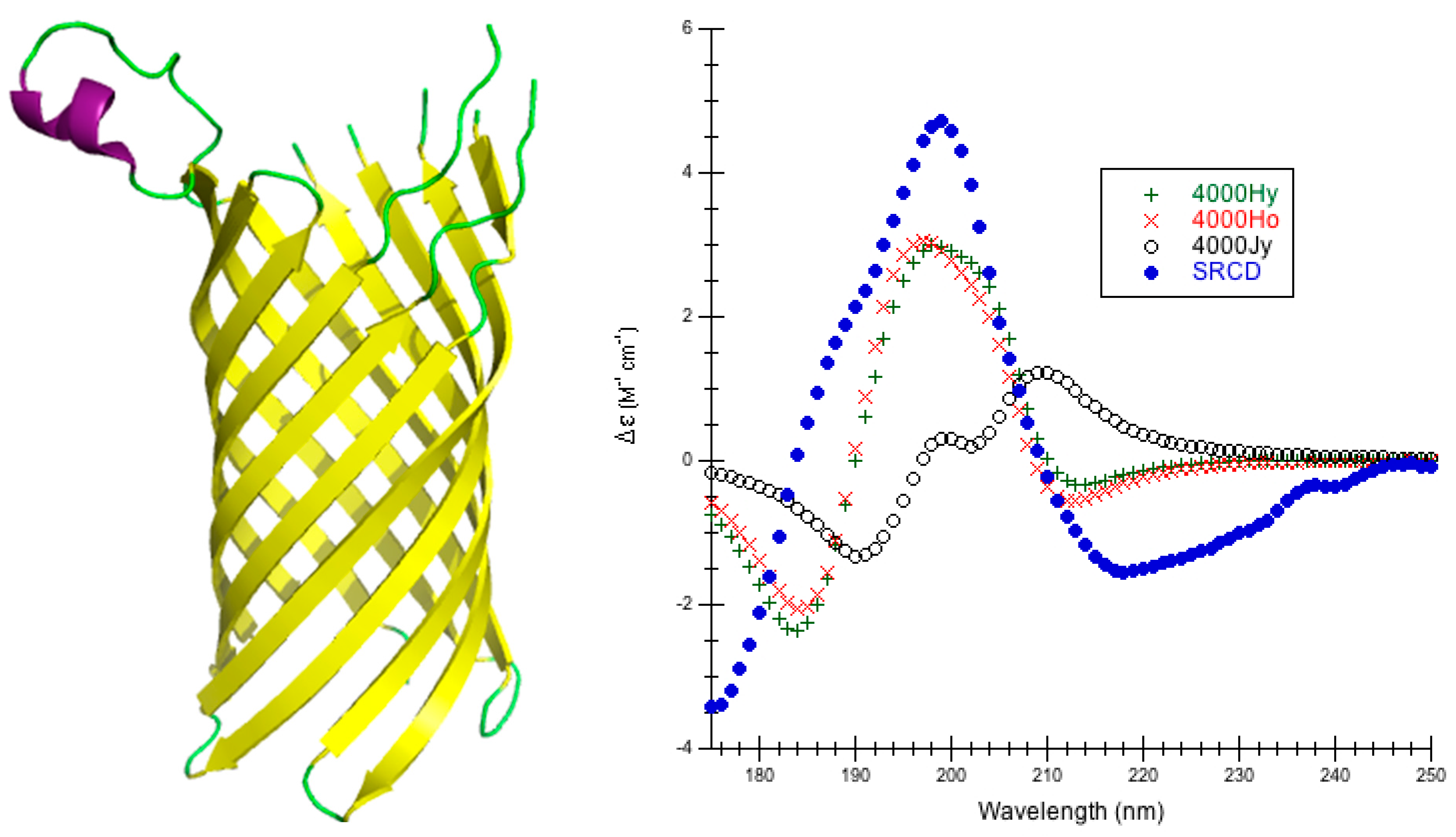
IJMS | Free Full-Text | Introducing DInaMo: A Package for Calculating Protein Circular Dichroism Using Classical Electromagnetic Theory

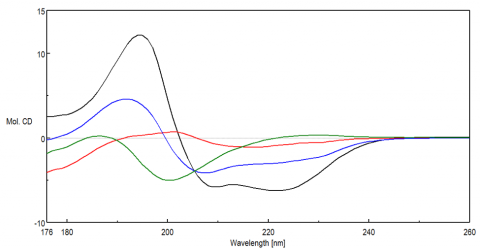
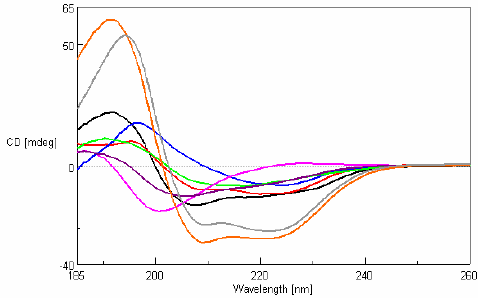
Analysis of changes in the secondary structure of full length Pol λ and it N-terminal deletion mutant proteins by far-UV CD spectroscopy.
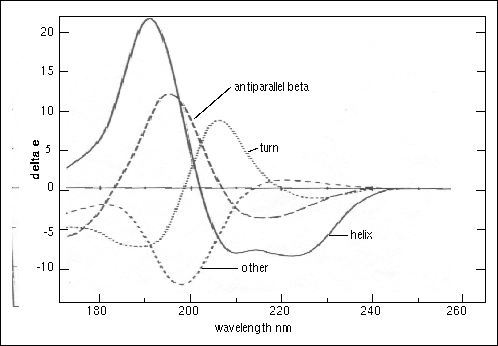
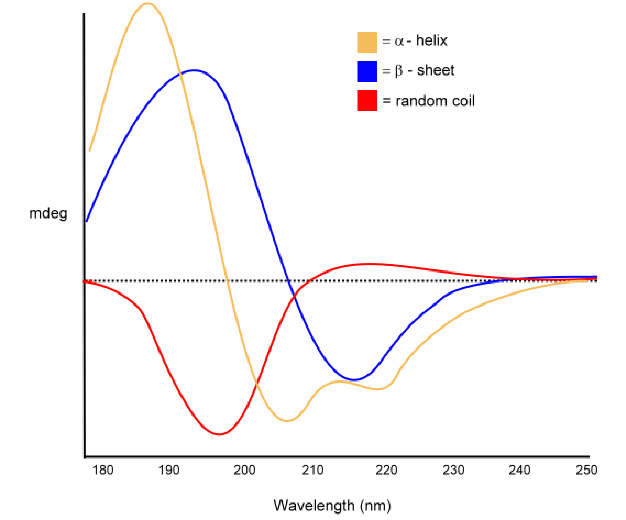
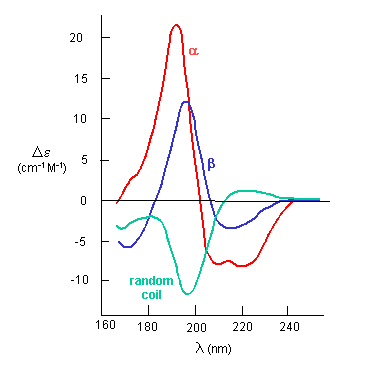
![Standard CD spectra redrawn from Corrêa et al.[5] Each of the three... | Download Scientific Diagram Standard CD spectra redrawn from Corrêa et al.[5] Each of the three... | Download Scientific Diagram](https://www.researchgate.net/publication/266950207/figure/fig14/AS:633005177597952@1527931603170/Standard-CD-spectra-redrawn-from-Correa-et-al5-Each-of-the-three-basic-secondary.png)
Standard CD spectra redrawn from Corrêa et al.[5] Each of the three... | Download Scientific Diagram
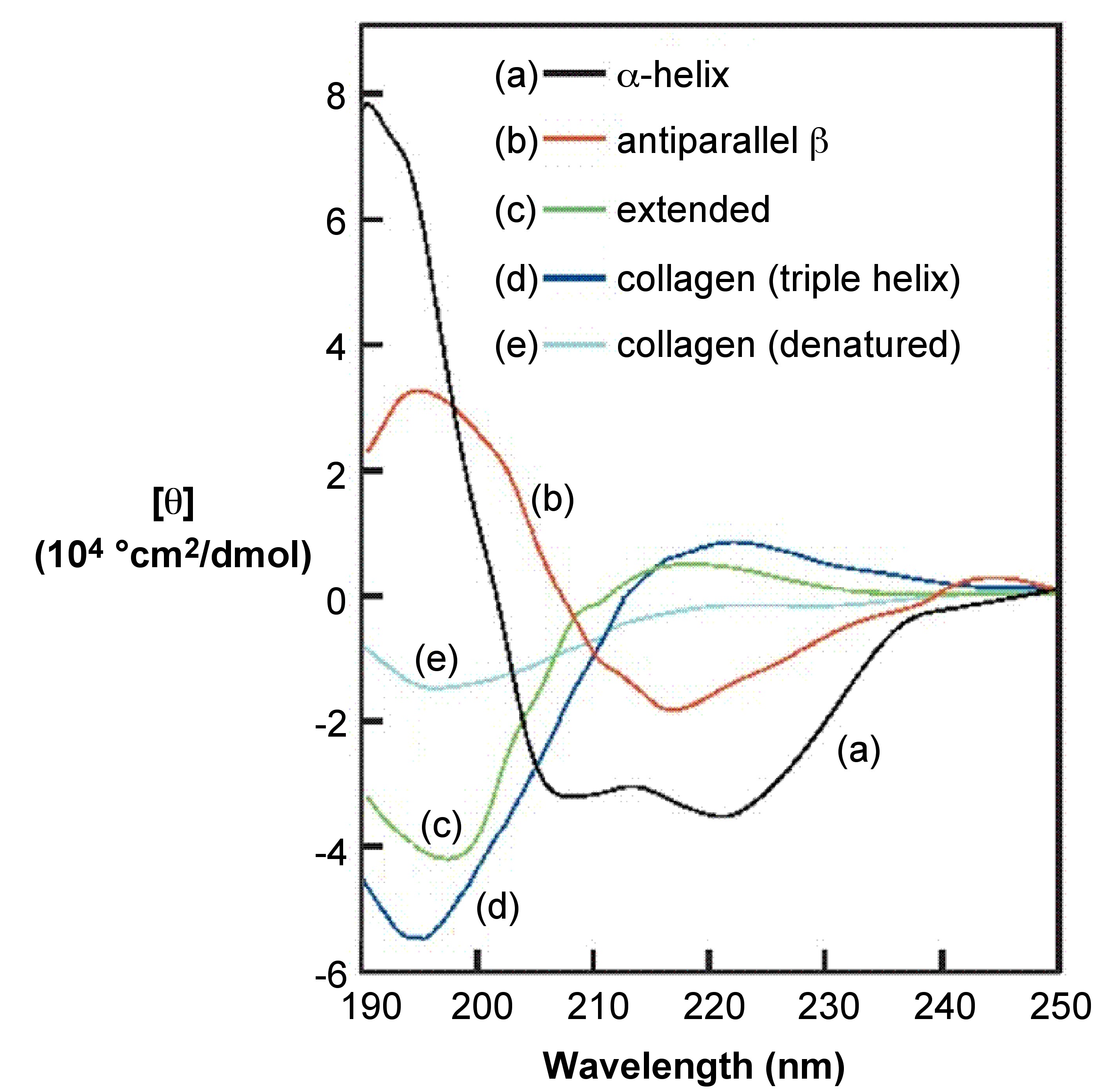
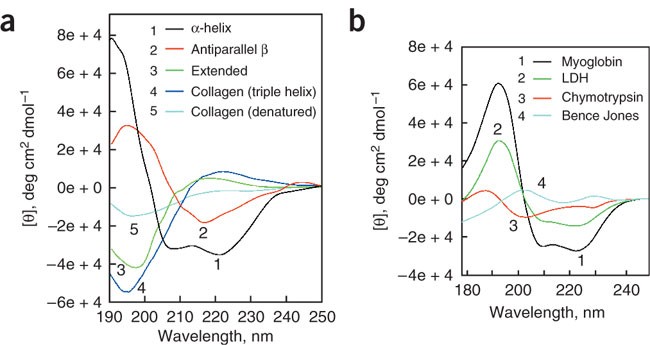
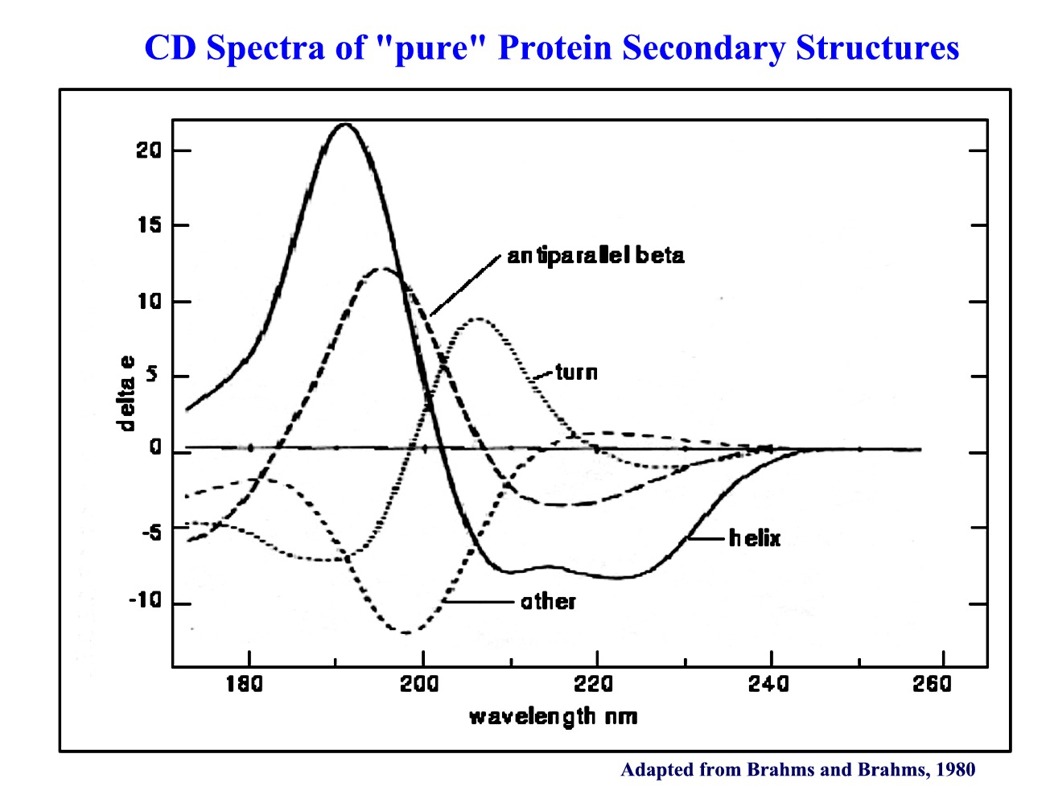
7.7: Circular Dichroism Spectroscopy and its Application for Determination of Secondary Structure of Optically Active Species - Chemistry LibreTexts

Secondary-Structure Analysis of Denatured Proteins by Vacuum-Ultraviolet Circular Dichroism Spectroscopy - ScienceDirect
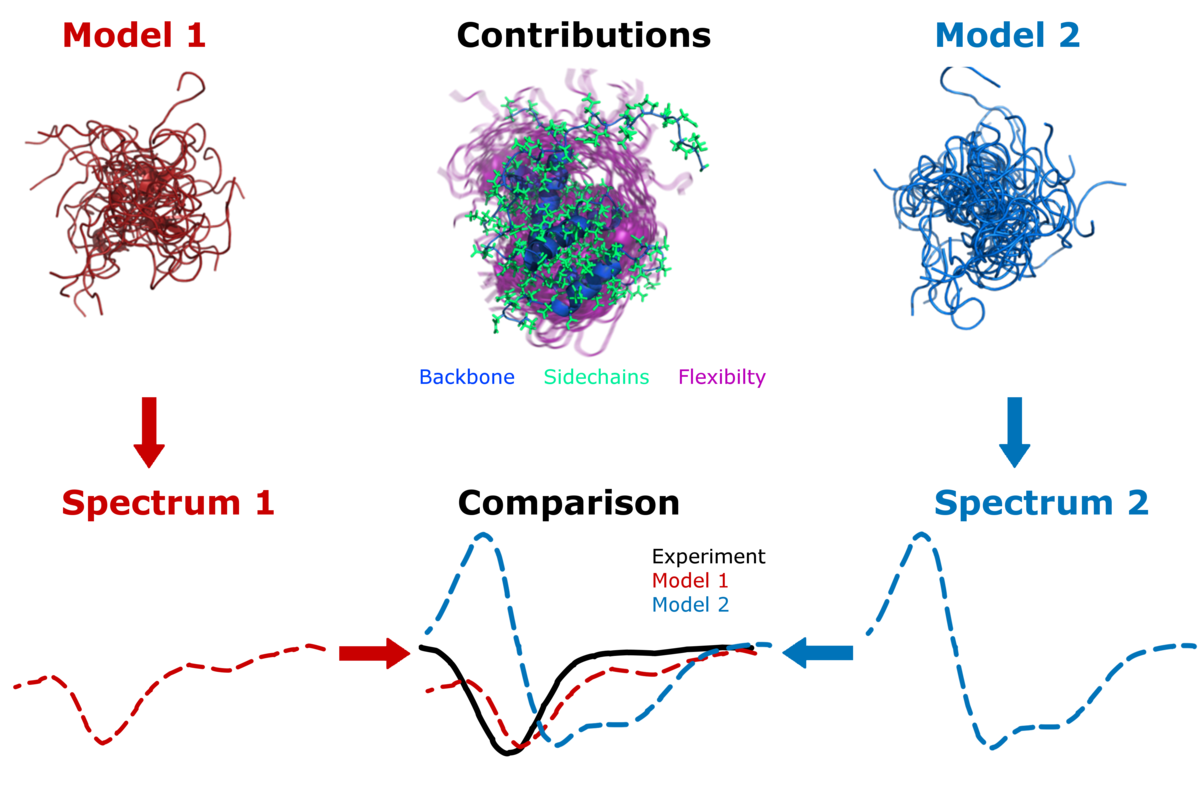
SESCA: Structure-based Empirical Spectrum Calculation Algorithm | Max Planck Institute for Multidisciplinary Sciences